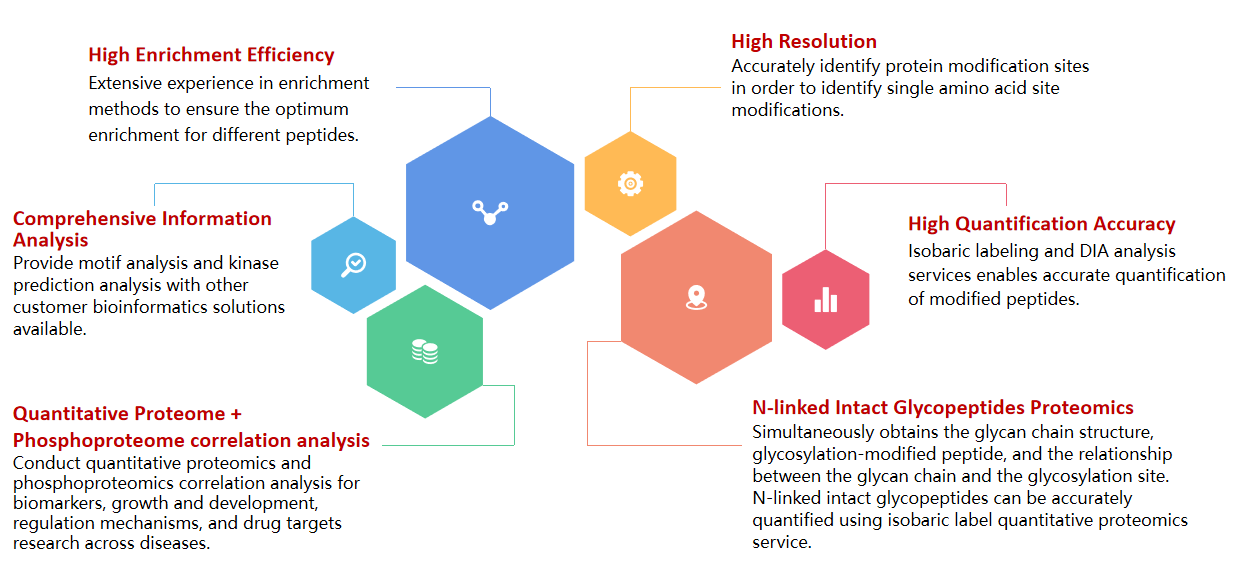
Service Description
PTM Proteomics
Post-translational modification (PTM) of proteins refers to the chemical changes proteins may undergo after translation. Common PTMs include Phosphorylation, Glycosylation, Acetylation and Ubiquitination. As a result, identifying and understanding PTMs is key for the study of cell biology and the treatment and prevention of disease.
BGI Genomics has extensive experience in the field of PTM Proteomics1,2 and has developed reliable workflows using market leading technologies and an advanced bioinformatics infrastructure.
Project Workflow
Phosphorylation:
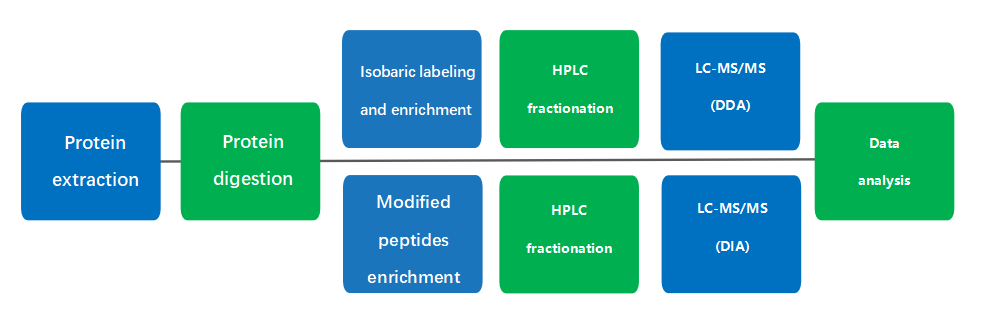
Acetylation/Ubiquitylation:
- Protein extraction
- Protein digestion
- Modified peptides enrichment
- HPLC fractionation
- LC-MS/MS (DDA)
- Data analysis
Glycosylaion:
- Protein extraction
- Protein digestion
- Modified peptides enrichment
- Remove glycan chains
- LC-MS/MS(DDA)
- Data analysis
- Protein extraction
- Protein digestion
- Isobaric labeling and enrichment
- HPLC fractionation
- LC-MS/MS
- Analysis by StrucGP
References
[1] Zhang, Y., et al., Evidence for Differential Glycosylation of Trophoblast Cell Types. Mol Cell Proteomics, 2016 Jun;15(6):1857-66. doi: 10.1074/mcp.M115.055798.
[2] Hao P L, et al., Correction of Errors in Tandem Mass Spectrum Extraction Enhances Phosphopeptide Identification. J. Proteome Res., 2013, 12, 12, 5548–5557. doi: 10.1021/pr4004486.
How to order
Mass Spectrometry Service Specification
Sample Preparation and Services
- Digestion performed using sequencing-grade trypsin
- Off-line sample fractionation using HPLC technology
- Each fraction analyzed using nano-flow LC-MS/MS
- TiO₂/ZrO₂ enrichment for phosphoproteomics; CST antibody enrichment for acetylated/ ubiquitylated proteins; MAX column enrichment for glycosylated proteins

Quality Standard
- Summary includes all methods and data analysis
- Reports provided in Excel or PDF format, RAW files available upon request

Turn Around Time
- Phosphorylation Turn Around Time from sample QC acceptance to data report delivery 3-5 weeks
- Glycosylation Turn Around Time from sample QC acceptance to data report delivery 4-5 weeks
- Acetylation/Ubiquitylation Turn Around Time from sample QC acceptance to data report delivery 3-5 weeks
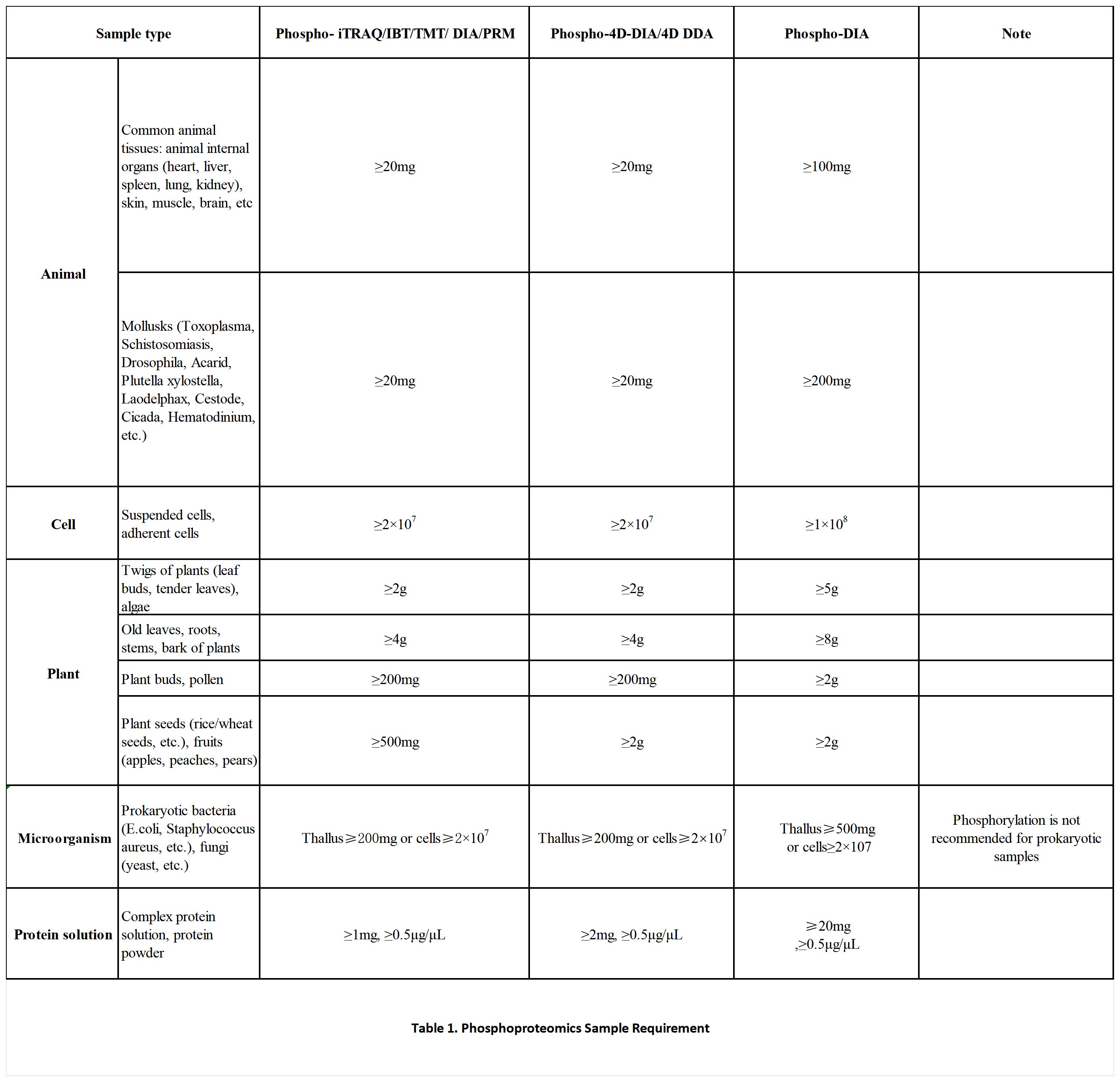
Sample Requirements
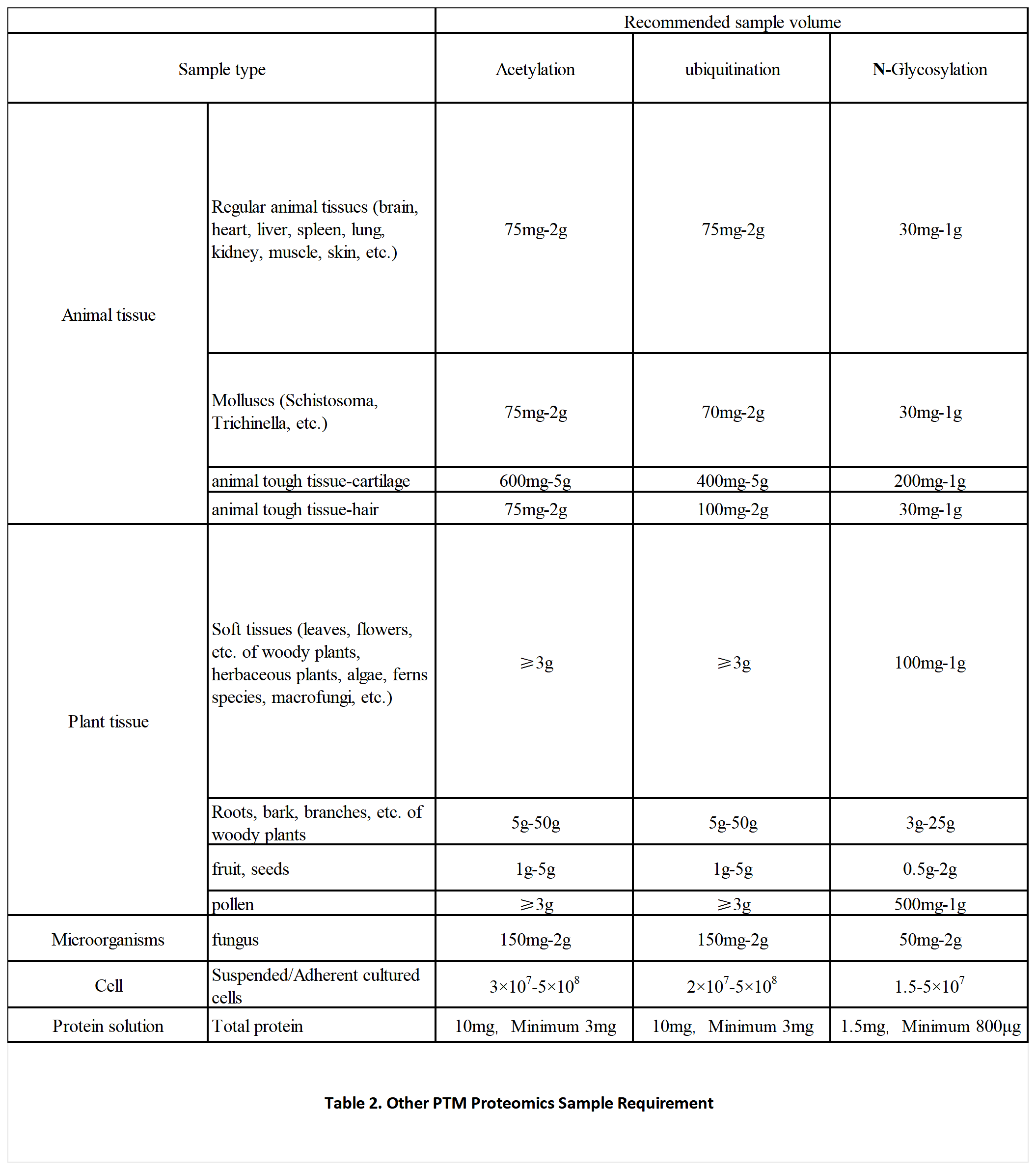
Data Analysis
Data analysis and validation performed with Mascot
Protein PTM identification
Protein PTM quantification
Modified protein function annotation
Differential modified protein function enrichment
Kinase prediction analysis
Motif distribution of PTM sites

